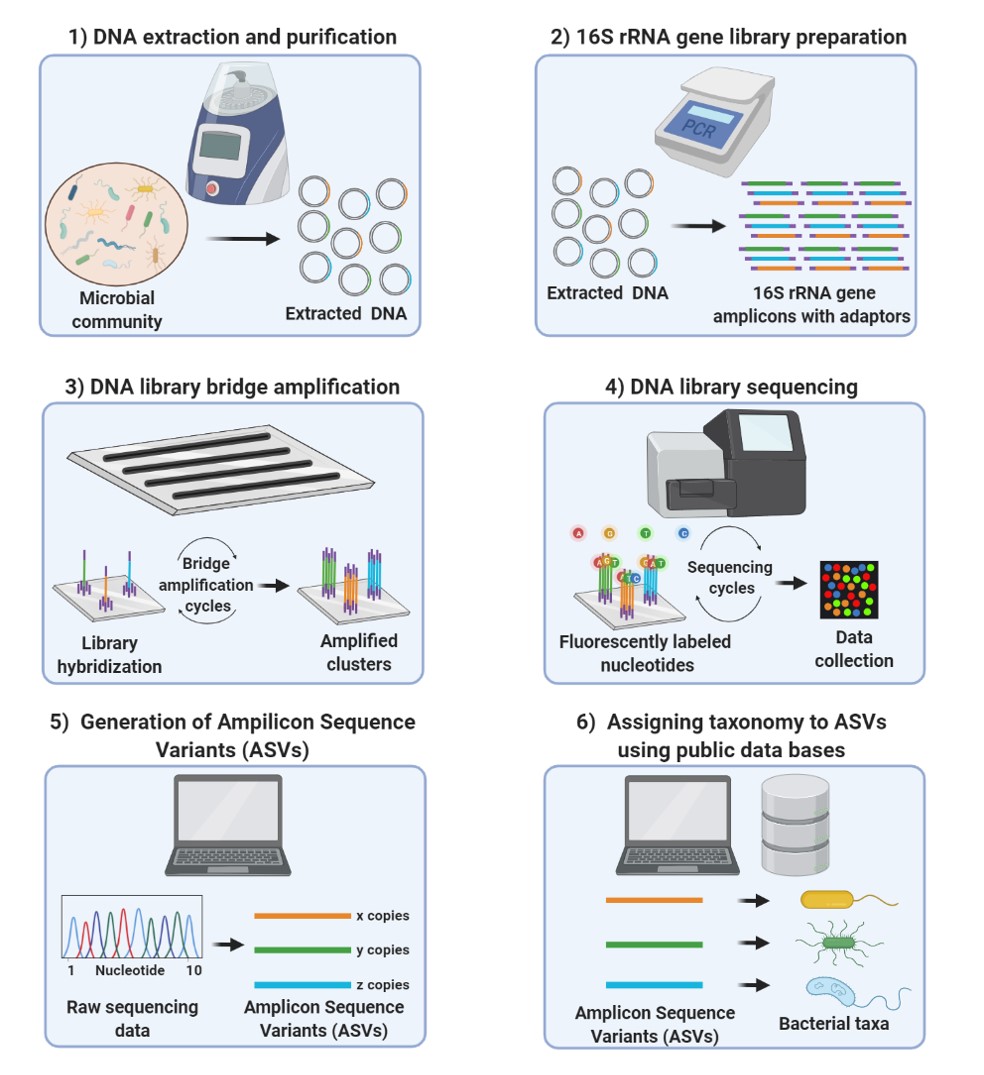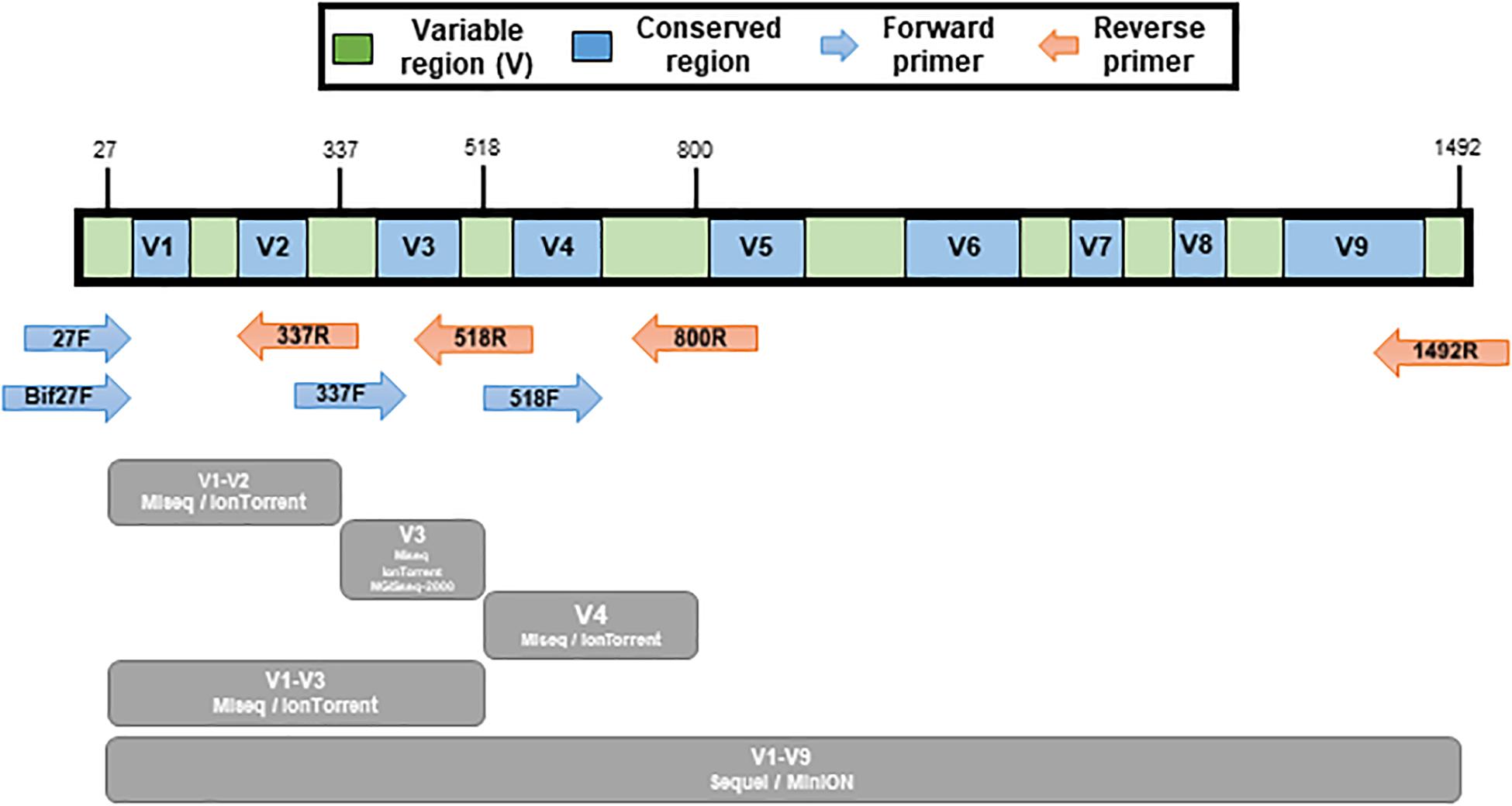
Frontiers | Comparison of 16S rRNA Gene Based Microbial Profiling Using Five Next-Generation Sequencers and Various Primers
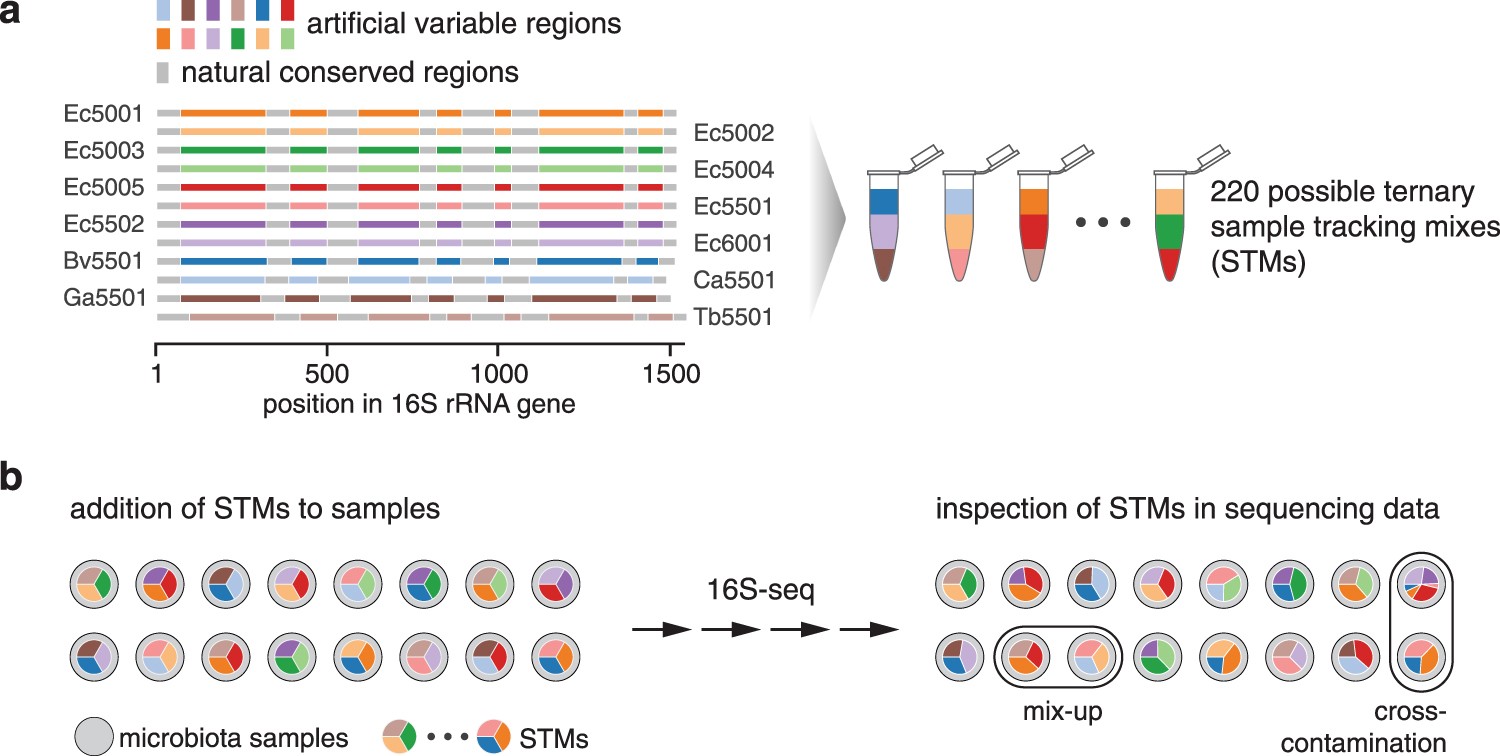
Sample tracking in microbiome community profiling assays using synthetic 16S rRNA gene spike-in controls | Scientific Reports
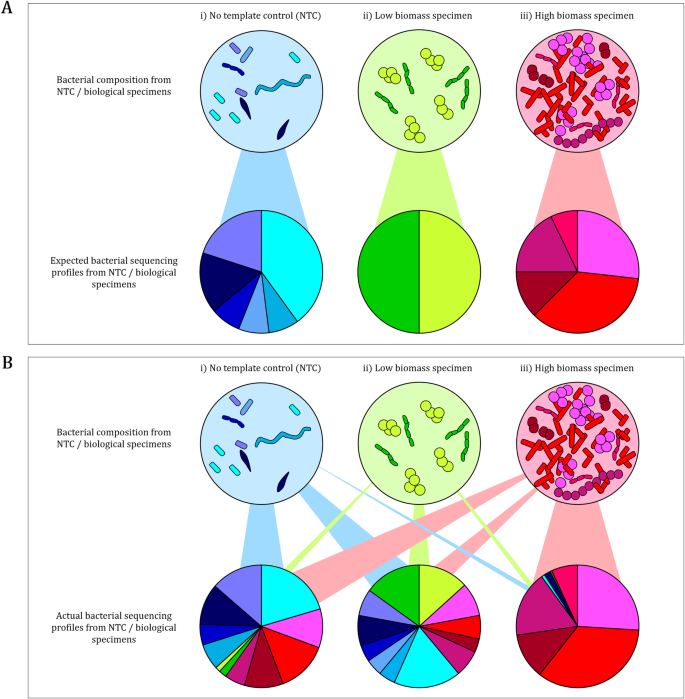
Optimizing 16S rRNA gene profile analysis from low biomass nasopharyngeal and induced sputum specimens | BMC Microbiology | Full Text

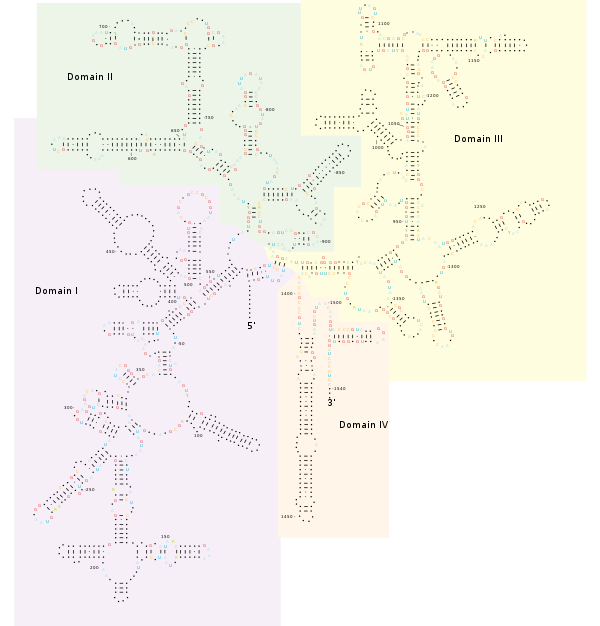
The schema of ribosome complex and 16S rRNA gene. The white and grey... | Download Scientific Diagram
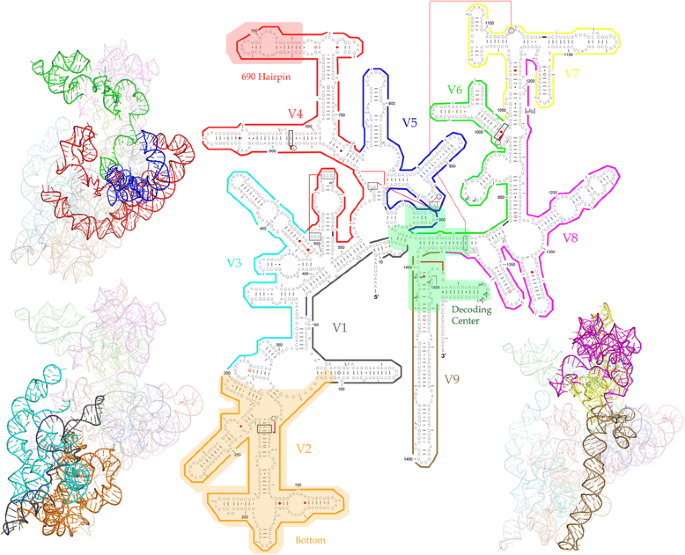
Sensitivity and correlation of hypervariable regions in 16S rRNA genes in phylogenetic analysis | BMC Bioinformatics | Full Text

Determining the most accurate 16S rRNA hypervariable region for taxonomic identification from respiratory samples | Scientific Reports

Uniting the classification of cultured and uncultured bacteria and archaea using 16S rRNA gene sequences | Nature Reviews Microbiology
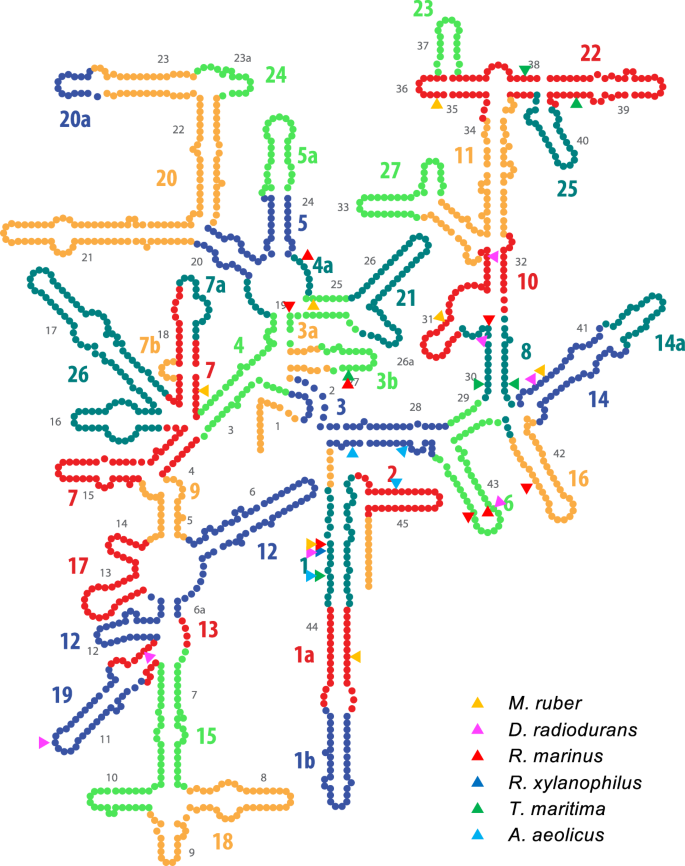
![De novo species identification using 16S rRNA gene nanopore sequencing [PeerJ] De novo species identification using 16S rRNA gene nanopore sequencing [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2020/10029/1/fig-1-full.png)