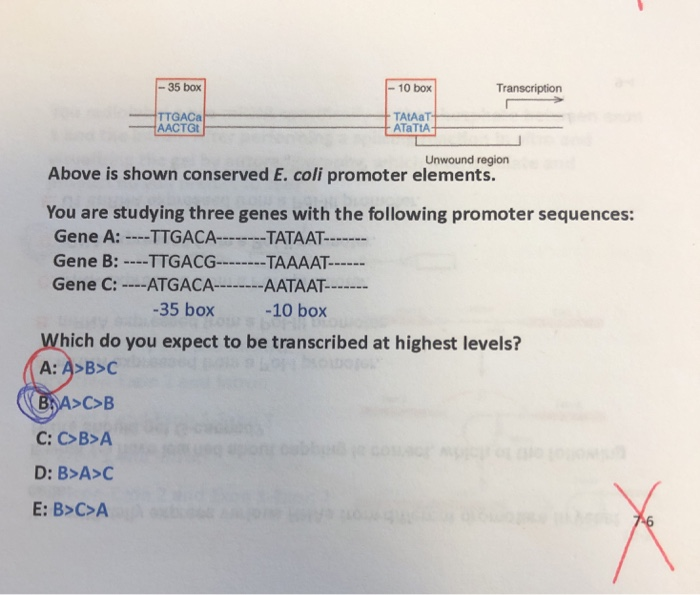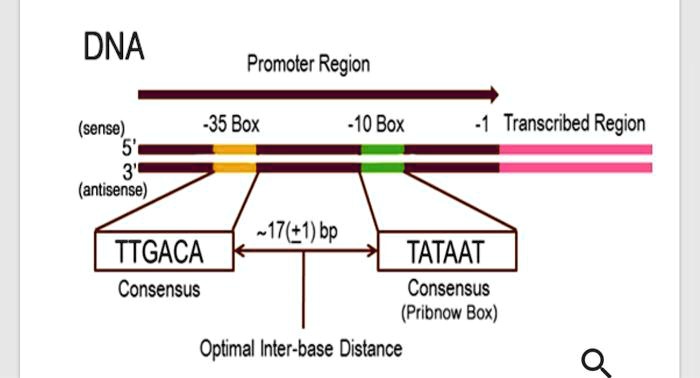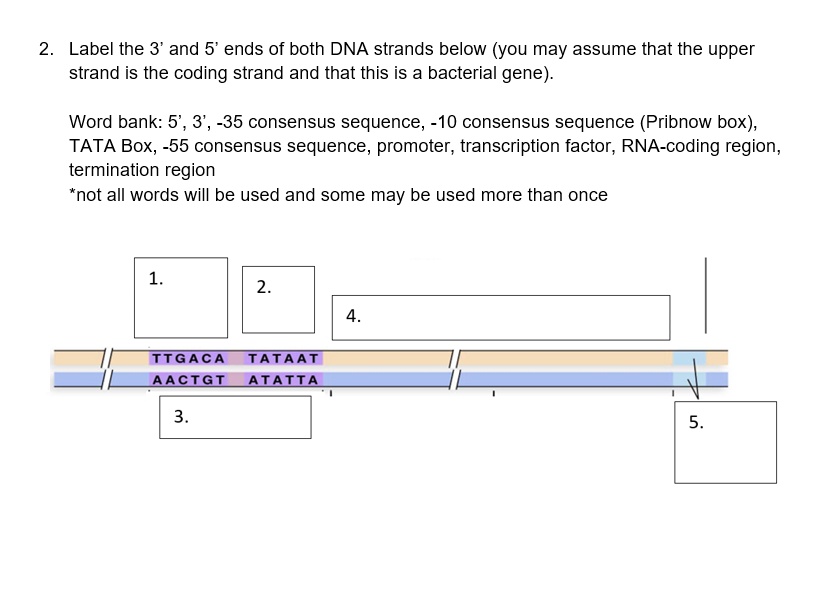
SOLVED: Label the 3' and 5' ends of both DNA strands below (you may assume that the upper strand is the coding strand and that this is a bacterial gene): Word bank:
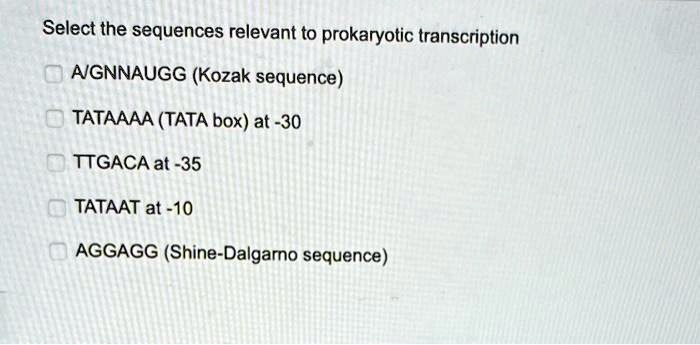

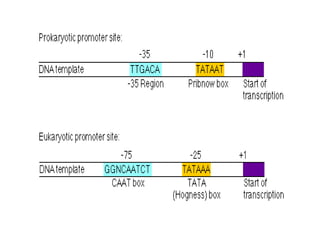
SOLVED: Texts: Select the sequences relevant to prokaryotic transcription: 1. A/GNNAUGG (Kozak sequence) 2. TATAAAA (TATA box at -30) 3. TTGACA at -35 4. TATAAT at -10 5. AGGAGG (Shine-Dalgarno sequence)